Project P03
P03 Human cytomegalovirus manipulation of HLA-E by US10
Dr. Anne Halenius
The human immune system can cover the presentation of a wide breadth of pathogen derived antigenic peptides owing to the three extraordinary polymorphic MHC-I genes loci each individual possesses. MHC-I molecules are expressed on the cell surface to communicate primarily with cytotoxic lymphocytes. While mainly the highly polymorphic classical MHC-I (HLA-A/B/C) present endogenous peptides to surveilling CD8+ T-cells, all MHC-I, including non-classical HLA-E/F/G, are able to interact with various inhibitory and/or activating NK cell receptors (NKR).
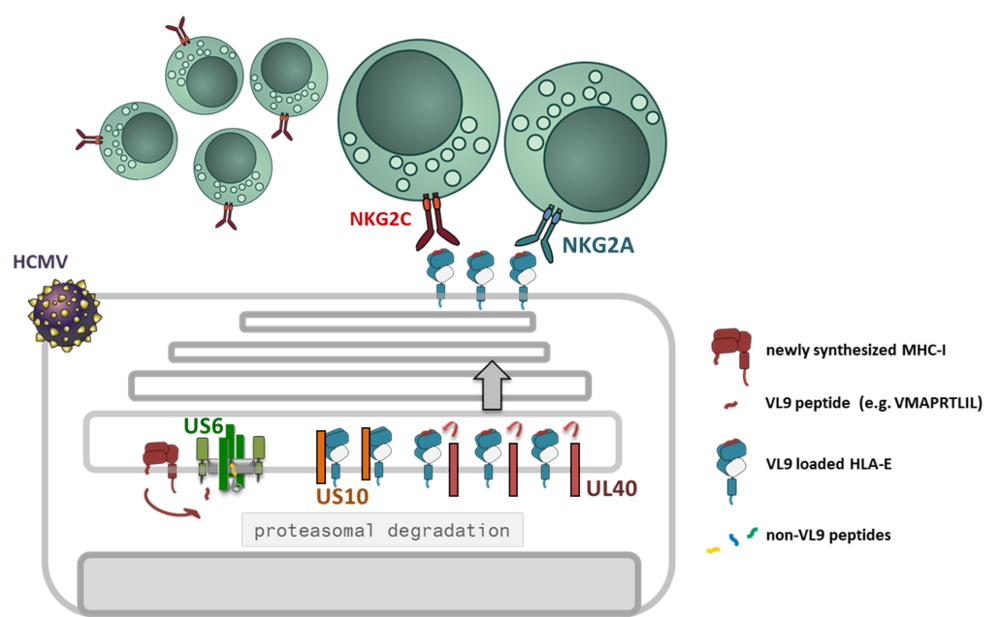
The unique infection biology of cytomegaloviruses has come along with the evolution of a large array of highly specialized immune evasive genes and reprogramming of multiple immune functions without compromising the human host [1]. HCMV controls distinct checkpoints of the MHC-I antigen presentation pathway by at least five gene products: US2 and US11 degrade MHC-I, US3 retains MHC-I inside the cells and US6 blocks the function of the peptide transporter TAP [2]. In addition, the viral miRNA miR-UL112-5p reduces the expression of ERAP1, a peptide trimming enzyme [3]. Thus, the control of CD8+ T-cell activation through manipulation of classical MHC-I is established and broadly documented. However, how HCMV might interfere with the MHC-I/NKR axis is not well understood.
Interestingly, a highly ADCC reactive NK cell population, also called memory-like NK cells (NKG2Cbright), has been found to expand uniquely in HCMV positive individuals [4, 5]. This NK cell subset exhibit a polarized KIR (killer cell immunoglobulin-like receptor) profile, with prominent expression of self-specific KIRs [6]. Expansion of memory-like NK cells in in vitro co-cultures with HCMV infected fibroblasts revealed an absolute dependency on HLA-E expression [7]. HLA-E is stabilized by MHC-I signal peptides (VL9) and engages in addition to the activating CD94/NKG2C NK cell receptor the inhibitory counterpart CD94/NKG2A. Two HCMV encoded proteins are known to affect HLA-E expression: since US6 blocks TAP function it deprives VL9 ligands from the ER, whereas UL40, which has pirated the VL9 peptide sequence into its signal peptide, loads HLA-E in a TAP-independent manner, i.e. in the presence of US6. Furthermore, we found that the glycoprotein US10 modulates HLA-E expression in HCMV infection.
In this project we will study the concerted molecular manipulation of HLA-E by known HCMV encoded immunoevasins and novel regulators and their effect on immune cell recognition of infected cells.
References
- Brodin P, Jojic V, Gao T, Bhattacharya S, Angel CJ, Furman D, et al. Variation in the human immune system is largely driven by non-heritable influences. Cell. 160:37-47 (2015)
- Halenius A, Gerke C, Hengel H. Classical and non-classical MHC I molecule manipulation by human cytomegalovirus: so many targets-but how many arrows in the quiver? Cellular & molecular immunology. 12:139-53 (2015)
- Romania P, Cifaldi L, Pignoloni B, Starc N, D'Alicandro V, Melaiu O, et al. Identification of a Genetic Variation in ERAP1 Aminopeptidase that Prevents Human Cytomegalovirus miR-UL112-5p-Mediated Immunoevasion. Cell reports. 20:846-53 (2017)
- Guma M, Angulo A, Vilches C, Gomez-Lozano N, Malats N, Lopez-Botet M. Imprint of human cytomegalovirus infection on the NK cell receptor repertoire. Blood. 104:3664-71 (2004)
- Schlums H, Cichocki F, Tesi B, Theorell J, Beziat V, Holmes TD, et al. Cytomegalovirus infection drives adaptive epigenetic diversification of NK cells with altered signaling and effector function. Immunity. 42:443-56 (2015)
- Beziat V, Liu LL, Malmberg JA, Ivarsson MA, Sohlberg E, Bjorklund AT, et al. NK cell responses to cytomegalovirus infection lead to stable imprints in the human KIR re